3D Embryo Cell Migration Simulator
A 3D simulation of Deep Forming Cell collective migration on the surface of a spherical zebrafish embryo during epiboly. Uses AER coordinate system with WebSocket streaming and Three.js visualization.
Business Context
During zebrafish gastrulation, the Enveloping Layer expands to engulf the yolk cell in a process called epiboly — one of the fundamental morphogenetic movements in vertebrate development. Deep Forming Cells ride this expansion, migrating collectively on the curved embryo surface. Simulating this process on a flat 2D plane introduces geometric distortions that compromise biological accuracy — the spherical geometry of the embryo is essential to understanding the migration dynamics.
Strategic Value
The simulation uses an AER (Azimuth-Elevation-Radius) coordinate system naturally suited to spherical geometry, with periodic wrapping at the azimuth and pole clamping at elevation. Cell velocity combines deterministic EVL coupling with Gaussian stochastic biological variability. Pairwise collision detection in angular space with symmetric push-apart resolution preserves momentum balance. Three.js visualization with orbit controls enables 3D embryo exploration. WebSocket streaming provides real-time simulation state. Developed at SCIAN-Lab, Universidad de Chile.
The Challenge
Simulating collective cell migration on a curved embryonic surface requires coordinate systems that handle spherical geometry — pole singularities, periodic wrapping — while maintaining physically meaningful collision detection and force resolution.
Our Approach
AER (Azimuth-Elevation-Radius) coordinate system with automatic wrapping at poles. EVL coupling provides deterministic base velocity; Gaussian stochastic term adds biological noise. Pairwise collision detection in angular space with symmetric push-apart resolution and sphere projection. WebSocket streaming for real-time 3D visualization.
Key Performance Indicators
| KPI | Baseline | Result | Impact |
|---|---|---|---|
| Geometry | 2D flat simulations | 3D spherical embryo surface | Biologically accurate geometry |
| Visualization | Offline rendering | Real-time Three.js WebGL | Interactive 3D exploration |
Architecture
sphersim
The Process
During zebrafish gastrulation, the Enveloping Layer (EVL) expands to engulf the yolk cell in a process called epiboly. Deep Forming Cells (DFCs) ride this expansion, migrating collectively on the curved embryo surface. This simulation captures that process on the actual spherical geometry of the embryo — not a flat approximation.
Spherical Coordinates
The AER (Azimuth-Elevation-Radius) coordinate system handles the geometry naturally. Azimuth wraps periodically (-π to π) — moving past one side means appearing on the other. Elevation clamps at the poles (-π/2 to π/2). Radius stays constant — cells are confined to the embryo surface.
Each cell’s velocity combines a deterministic component from EVL coupling (base_velocity = [0, -(π/2)·evl_speed, 0] — the expanding EVL drags cells vegetally) with a Gaussian stochastic term adding biological variability. Collision detection operates in angular space with symmetric push-apart resolution: overlapping cells are displaced equally in opposite directions, then projected back onto the sphere.
Three.js visualization with orbit controls lets researchers rotate, zoom, and inspect the 3D embryo from any angle. WebSocket streaming provides real-time simulation state. Play/Pause/Step controls enable frame-by-frame inspection of migration dynamics.
Technology Stack
Application Screenshots
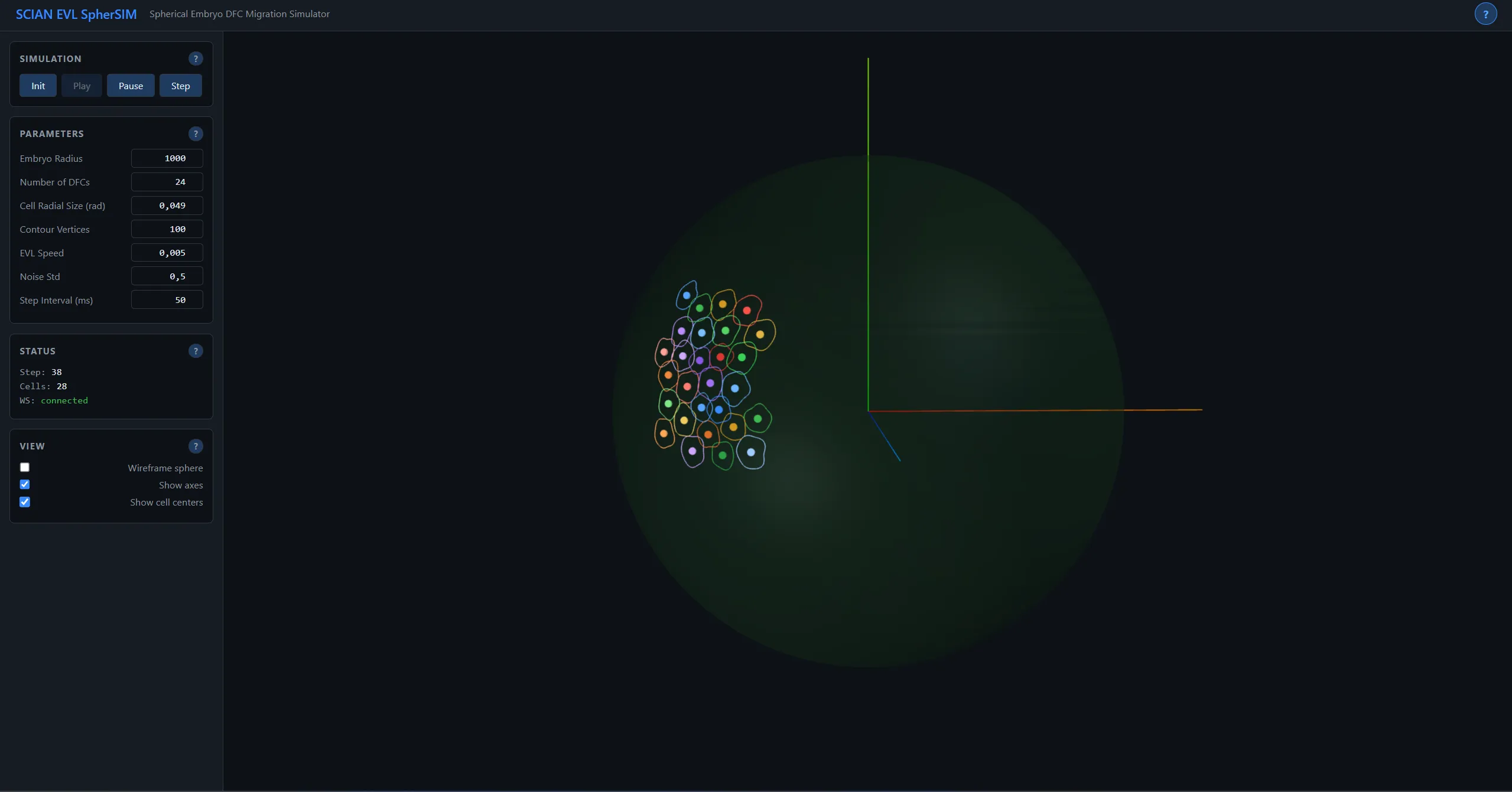
Technical Diagrams
evl spring coupling
sphersim coordinates
sphersim epiboly
sphersim model